Notes
This page describes some technical details of our Peptide Calculator.
Calculating the molecular weight
The molecular weight M of a peptide may be estimated by calculating
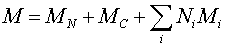
where Ni are the number, and Mi
the average residue molecular weights, of the amino acids. MN + MC are
added to the total in order to account for the termini: H at the N-terminus and OH at the C-terminus. Of course,
if the termini are modified, these additions are replaced by those of the modifiers.
The resulting molecular weight depends on what values the algorithm uses. Some of the values Innovagen's
Peptide Calculator uses are:
| A, Ala | 71.07793 | C, Cys | 103.1454 | D, Asp | 115.0873 | E, Glu | 129.1139 |
| F, Phe | 147.1734 | G, Gly | 57.05138 | H, His | 137.1394 | I, Ile | 113.1576 |
| K, Lys | 128.1724 | L, Leu | 113.1576 | M, Met | 131.1985 | N, Asn | 114.1028 |
| P, Pro | 97.11508 | Q, Gln | 128.1293 | R, Arg | 156.1861 | S, Ser | 87.07733 |
| T, Thr | 101.1039 | V, Val | 99.13103 | W, Trp | 186.2095 | Y, Tyr | 163.1728 |
| H | 1.00797 | OH | 17.00738 | phos-Ser | 167.0573 | ||
| Acetyl | 43.04453 | Amide | 16.0228 | phos-Thr | 181.0838 | ||
| Biotin | 227.3056 | phos-Tyr | 243.1528 | ||||
Calculating the extinction coefficient
The molar extinction coefficient e at 280 nm of a peptide can be estimated by calculating
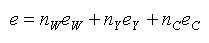
where nW, nY, and nC are the number of Tryptophans (W), Tyrosines (Y) and Cystines (i.e. disulphide bonds, but here denoted C) in the sequence. The molar extinction coefficients used in Innovagen's Peptide Calculator are:
| eW = 5690 M-1cm-1 | eY = 1280 M-1cm-1 | eC = 120 M-1cm-1 | |
| Gill and von Hippel | |||
Note: Your peptide may contain non-natural amino acids, modifications and labels. None of these are considered, although they may very well add to the extinction coefficient.
Calculating the net charge
The net charge Z of a peptide at a certain pH can be estimated by calculating
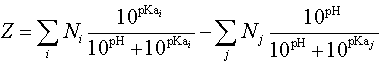
where Ni are the number, and pKai
the pKa values, of the N-terminus and the side chains of Arginine, Lysine,
and Histidine. The j-index pertain to the C-terminus and the Aspartic
Acid, Glutamic Acid, Cysteine, Tyrosine amino acids.
Innovagen's Peptide Property Calculator calculates the net
charge for all pH values of 0.1 to 14 in increments of 0.1, and plots these producing a titration curve.
This algorithm has its limitations, some of which are:
- The residues are assumed to be independent of each other.
- Only free terminii and the 20 naturally occuring amino acids and their D-forms are considered, all others are ignored.
- The resulting net charge depends on what pKa values the algorithm uses. Innovagen's Peptide Property Calculator uses the values taken from the CRC Handbook of Chemistry and Physics, 87th edition.